ZULE’s Data & GitHub Crash Course
2024-01-30
Part I: Prep Work
Learning Goals
My goal for this workshop is to give everyone the tools to:
- Confidently start a project in R
- Manage files in a way that is reproducible and easy to understand
- Allow people to document history/progress on their projects
- Know one approach to publicly archiving projects
Software Installation
- CHECK-IN: does everyone have everything working/installed?
- Absolutely the hardest part of this workshop
- Thank you for doing homework!!
- If you have technical issues throughout this presentation - raise your hand and we will either work through together or Kayleigh will help you troubleshoot
Transparent Workflows
Ensuring that your workflow is transparent is important for:
Past/Current/Future You
ZULE Lab
Collaborators
Other grad students
Scientific Community
PUBLIC
Part II: R & RStudio
Project Management in R
Good file structure is important because it 1
- Ensures the integrity of your data
- Makes it easier to share your code with people
- Makes it easier to upload your code/data with manuscript submission
- Makes it easier to come back after a break
File Management for R
Best practices include (but are not limited to) 1
- Use an R Project file so that your project is easily shareable
- Always treat raw data as read-only
- Store cleaned data in a separate folder (or distinguish clearly)
- Treat output as disposable - you should always be able to re-generate with script
- Have separate function and figure scripts
Cleaning Data in R
- Reproducible 1
- Open-source and cross-platform
- Reliable & clear
- High-quality graphics
- Great community & resources
- Scales with datasets
- Steep learning curve with a high payoff
Cleaning Data in R
There are some tasks that do not need to be “as reproducible” (e.g., fixing typos) - these can be done in OpenRefine.
In general if you are:
Combining data sources
Making decisions about the data itself (e.g., removing or adding data)
Performing calculations
Renaming things
Do this in R (you will be grateful later!)
“Tidy” (or clean) Data:
- Framework for how data should be formatted for easy and efficient data cleaning created by Hadley Wickham
- Underpinnings of
tidyversepackages (e.g. ggplot2)
- Underpinnings of
Principles:
- Each variable forms a column
- Each observation forms a row
- Each type of observational unit forms a table
Basic File Structure

Let’s Make a Project
Let’s set up a new project, using RProjects
Add input, output, script, and figure folders
(I recommend you have a place on your computer dedicated to this)
Part II: GitHub
GitHub & Version Control

Piled Higher and Deeper
GitHub & Version Control
GitHub is a website-software that documents your progress on a project and allows you to do version control
- aka it takes snapshots of your progress across time so nothing gets lost
If you save rough drafts of your writing as you go along - that is version control
Really useful for when you want to go back/change your mind/re-run a test/etc.
Facilitates peace of mind + reproducible science + collaboration/sharing
Project Workflow with Git
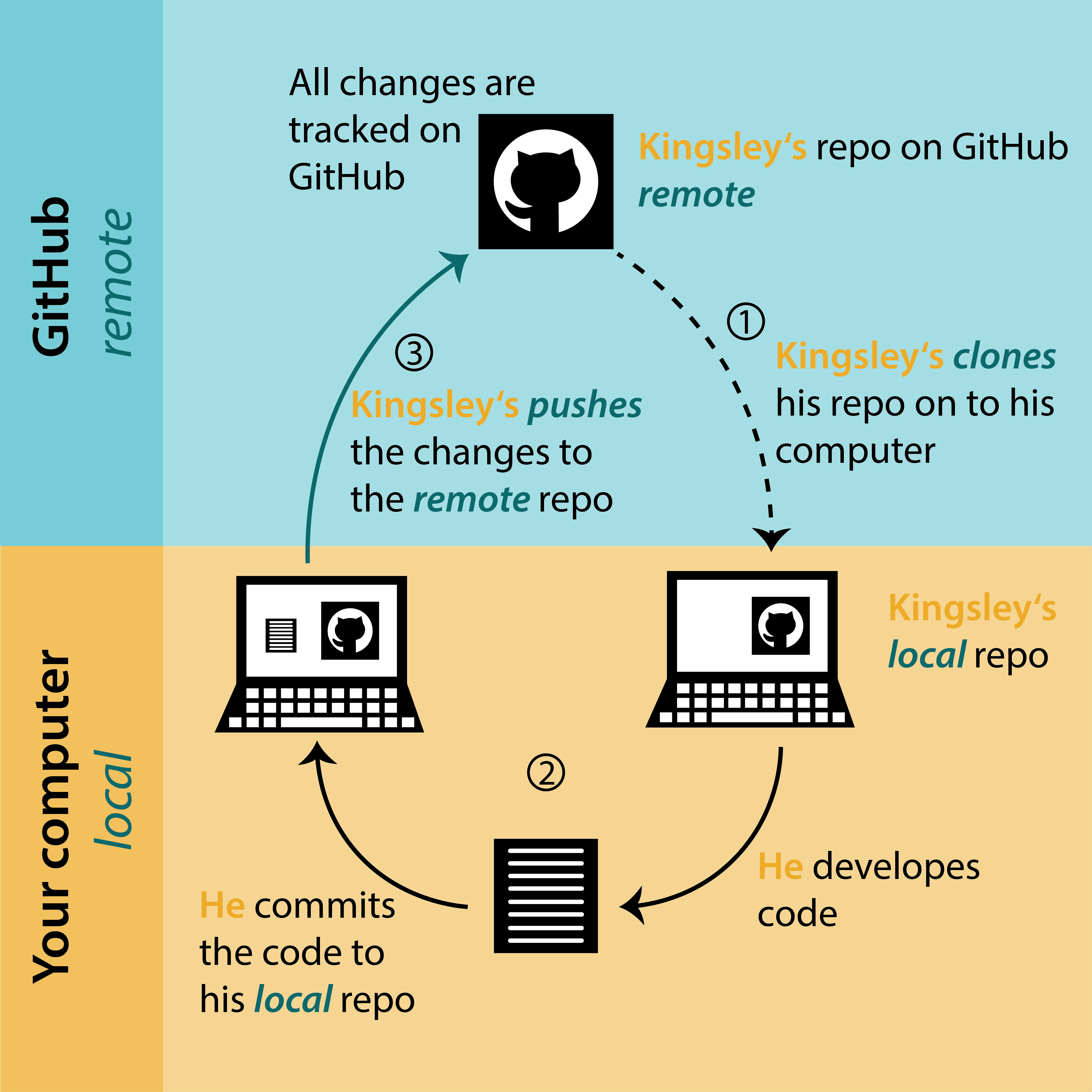
biost@ts Git Tutorial
Project Workflow with Git + Others
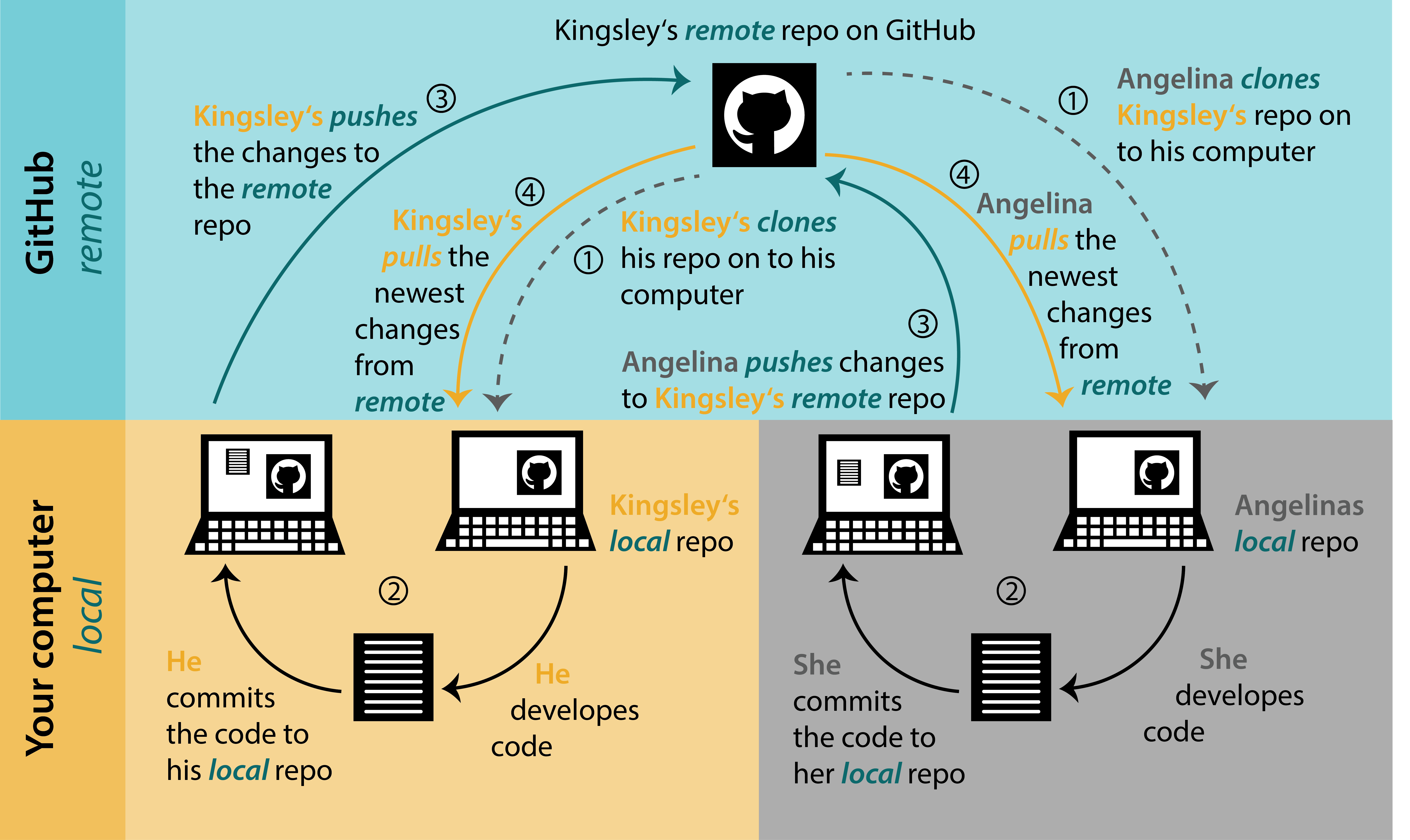
biost@ts Git Tutorial
The Basics of GitHub
- 5 basic jargon terms you need to know to use GitHub:
- Repository/repo: your project
- Clone: make a local copy of your project
- Commit: describe and commit to any changes you’ve made
- Push: send your changes to your online repo
- Pull: incorporate any changes to your local repo
- (BONUS branch: a side project)
- We will do all these things today!
Let’s Make a Repository!

Let’s Make a Repository!

ZULE’s GitHub has lots of repositories (including examples) if you are looking for inspiration for folder organization, ReadMe documentation, metadata, etc.
Let’s Make a Repository!

Cloning (Download An Existing Directory)

Committing
GitHub tracks the changes you make to your repository on your computer
After making changes, you have to select, describe, and commit them
Committing

Pushing
After committing, you push your changes to your remote repository

Pulling (Collaboration Station)
If you are collaborating on a project, where multiple people are contributing, make sure you pull from the remote repository before starting your work
Same button as push (ctrl + shift + P)
Part IV: Archiving Data
Lab Archiving
Archiving your project in the lab requires 4 things:
- Paper/thesis
- Clean data
- Metadata
- Code
These things can be organized however you’d like, as long as they are easily understood by someone after you are gone.
Projects need to be added to the lab computer, under the D: drive, in the Lab_Alumni folder
Why Using Git is not Archiving
Does not have a DOI, so does not point to a specific moment in time
Can be changed continuously
Not dedicated to longevity
Can import GitHub repository to a true data archive
Public Archiving
Zenodo is a great option for archiving data
Easily links to GitHub repositories
Preserves file structures
Can be updated after reviews/changes with a new DOI
FREE
Other options include Dryad, figshare, and more topic-specific archives (e.g., GenBank)
As always, use what works for you
Zenodo
To connect and archive your code/data with Zenodo from GitHub, there are three main steps
- Link your GitHub to your Zenodo account, and toggle “On” for your repository
- Make a release of the project on GitHub
- Obtain DOI and project page from Zenodo
(see an example workthrough here)
NOTE: you do not need to use Git to use Zenodo, you can also upload local files
The Ultimate Combo Deal


Resources
This workshop - including examples & code can all be found here and formatted slides are here
Software Carpentry: R for Reproducible Scientific Analysis & Version Control with git
Data Carpentry: Data Analysis & Visualization in R for Ecologists & Data Organization in Spreadsheets for Ecologists
biost@ts: Version Control with Git and GitHub
Happy Git: happygitwithr
University of Bergen: Open Access to Research Data
Resources
Smart People I Know: Dr. Christie Bahlai’s Reproducible Quantitative Methods Course & Wildlife Ecology & Evolution Lab’s Guide by Alec Robitaille & Val Lucet’s Git Workshop
PLUS: check out our zup “stats” thread - lots of helpful resources! AND ASK YOUR LABMATES!!!!